We’re downright exhilarated to announce the release of rsample 1.1.0. The rsample package makes it easy to create resamples for estimating distributions and assessing model performance.
You can install it from CRAN with:
install.packages("rsample")This blog post will walk through some of the highlights from this newest release. You can see a full list of changes in the release notes.
Grouped resampling
By far and away the biggest addition in this version of rsample is the set of new functions for grouped resampling. Grouped resampling is a form of resampling where observations need to be assigned to the analysis or assessment sets as a “group”, not split between the two. This is a common need when some of your data is more closely related than would be expected under random chance: for instance, when taking multiple measurements of a single patient over time, or when your data is geographically clustered into distinct “locations” like different neighborhoods.
The rsample package has supported grouped v-fold cross-validation for a few years, through the
group_vfold_cv() function:
library(purrr)
library(rsample)
data(ames, package = "modeldata")
resample <- group_vfold_cv(ames, group = Neighborhood, v = 2)
resample$splits %>%
map_lgl(function(x) {
any(assessment(x)$Neighborhood %in% analysis(x)$Neighborhood)
}
)
#> [1] FALSE FALSErsample 1.1.0 extends this support by adding four new functions for grouped resampling. The new functions
group_bootstraps(),
group_mc_cv(),
group_validation_split(), and
group_initial_split() all work like their ungrouped versions, but let you specify a grouping column to make sure related observations are all assigned to the same sets:
# Bootstrap resampling with replacement:
group_bootstraps(ames, Neighborhood, times = 1)
#> # Group bootstrap sampling
#> # A tibble: 1 × 2
#> splits id
#> <list> <chr>
#> 1 <split [3050/1225]> Bootstrap1
# Random resampling without replacement:
group_mc_cv(ames, Neighborhood, times = 1)
#> # Group Monte Carlo cross-validation (0.75/0.25) with 1 resamples
#> # A tibble: 1 × 2
#> splits id
#> <list> <chr>
#> 1 <split [2198/732]> Resample1
# Data splitting to create a validation set:
group_validation_split(ames, Neighborhood)
#> # Group Validation Set Split (0.75/0.25)
#> # A tibble: 1 × 2
#> splits id
#> <list> <chr>
#> 1 <split [2201/729]> validation
# Data splitting to create an initial training/testing split:
group_initial_split(ames, Neighborhood)
#> <Training/Testing/Total>
#> <2162/768/2930>These functions all target assigning a certain proportion of your data to the assessment fold. Hitting that target can be tricky when your groups aren’t all the same size, however. To work around this, these new functions create a list of all the groups in your data, randomly reshuffle it, and then select the first n groups in the list that results in splitting the data as close to that proportion as possible. The net effect of this on users is that your analysis and assessment folds won’t always be precisely the size you’re targeting (particularly if you have a few large groups), but all data in a single group will always be entirely assigned to the same set and the splits will be entirely randomly created.
The other big change to grouped resampling comes as a new argument to
group_vfold_cv(). By default,
group_vfold_cv() assigns roughly the same number of groups to each of your folds, so you wind up with the same number of patients, or neighborhoods, or whatever else you’re grouping by in each assessment set. The new balance argument lets you instead assign roughly the same number of rows to each fold instead, if you set balance = observations:
group_vfold_cv(ames, Neighborhood, balance = "observations")
#> # Group 28-fold cross-validation
#> # A tibble: 28 × 2
#> splits id
#> <list> <chr>
#> 1 <split [2928/2]> Resample01
#> 2 <split [2922/8]> Resample02
#> 3 <split [2907/23]> Resample03
#> 4 <split [2736/194]> Resample04
#> 5 <split [2886/44]> Resample05
#> 6 <split [2893/37]> Resample06
#> 7 <split [2929/1]> Resample07
#> 8 <split [2663/267]> Resample08
#> 9 <split [2805/125]> Resample09
#> 10 <split [2837/93]> Resample10
#> # … with 18 more rows
#> # ℹ Use `print(n = ...)` to see more rowsThis approach works in a similar way to the new grouped resampling functions, attempting to assign roughly 1 / v of your data to each fold. When working with unbalanced groups, this can result in much more even assignments of data to each fold:
library(ggplot2)
library(dplyr)
analysis_sd <- function(v, balance) {
group_vfold_cv(
ames,
Neighborhood,
v,
balance = balance
)$splits %>%
purrr::map_dbl(~ nrow(analysis(.x))) %>%
sd()
}
resample <- tidyr::crossing(
idx = seq_len(100),
v = c(2, 5, 10, 15),
balance = c("groups", "observations")
)
resample %>%
mutate(sd = purrr::pmap_dbl(
list(v, balance),
analysis_sd
)) %>%
ggplot(aes(sd, fill = balance)) +
geom_histogram(alpha = 0.6, color = "black", size = 0.3) +
facet_wrap(~ v) +
theme_minimal() +
labs(title = "sd() of nrow(analysis) by balance method")
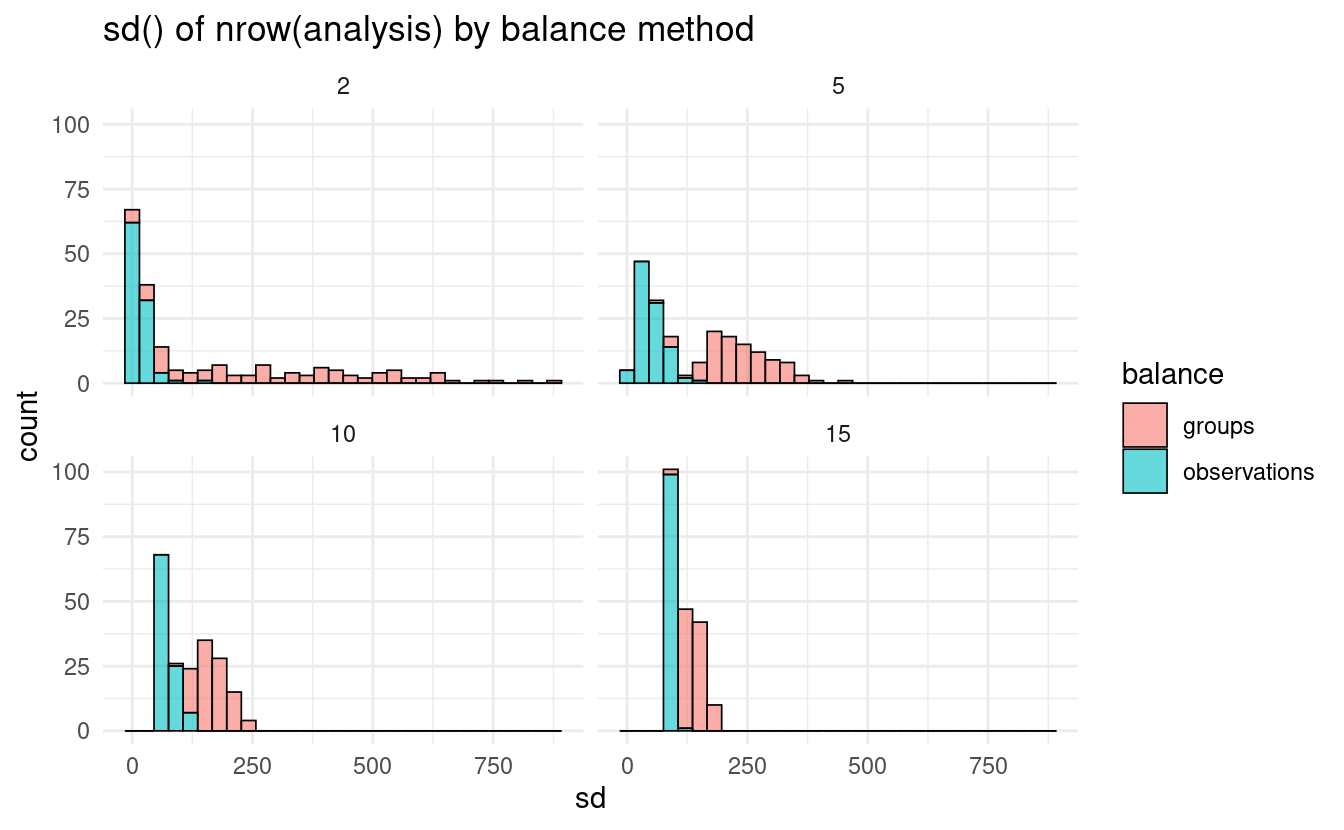
Right now, these grouping functions don’t support stratification. If you have thoughts on how you’d expect stratification to work with grouping, or have an example of how another implementation has handled it, let us know on GitHub!
Other improvements
This release also adds a few new utility functions to make it easier to work with the rsets produced by rsample functions.
For instance, the new
reshuffle_rset() will re-generate an rset, using the same arguments as were used to originally create it, but with the current random seed:
set.seed(123)
resample <- vfold_cv(mtcars)
resample$splits[[1]] %>%
analysis() %>%
head()
#> mpg cyl disp hp drat wt qsec vs am gear carb
#> Mazda RX4 Wag 21.0 6 160 110 3.90 2.875 17.02 0 1 4 4
#> Datsun 710 22.8 4 108 93 3.85 2.320 18.61 1 1 4 1
#> Hornet 4 Drive 21.4 6 258 110 3.08 3.215 19.44 1 0 3 1
#> Hornet Sportabout 18.7 8 360 175 3.15 3.440 17.02 0 0 3 2
#> Valiant 18.1 6 225 105 2.76 3.460 20.22 1 0 3 1
#> Duster 360 14.3 8 360 245 3.21 3.570 15.84 0 0 3 4
resample <- reshuffle_rset(resample)
resample$splits[[1]] %>%
analysis() %>%
head()
#> mpg cyl disp hp drat wt qsec vs am gear carb
#> Mazda RX4 21.0 6 160 110 3.90 2.620 16.46 0 1 4 4
#> Mazda RX4 Wag 21.0 6 160 110 3.90 2.875 17.02 0 1 4 4
#> Datsun 710 22.8 4 108 93 3.85 2.320 18.61 1 1 4 1
#> Hornet 4 Drive 21.4 6 258 110 3.08 3.215 19.44 1 0 3 1
#> Hornet Sportabout 18.7 8 360 175 3.15 3.440 17.02 0 0 3 2
#> Duster 360 14.3 8 360 245 3.21 3.570 15.84 0 0 3 4This works with repeated cross-validation, stratification, grouping – anything you did originally should be preserved when reshuffling the rset.
Additionally, the new
reverse_splits() function will “swap” the assessment and analysis folds of any rsplit or rset object:
resample <- initial_split(mtcars)
resample
#> <Training/Testing/Total>
#> <24/8/32>
reverse_splits(resample)
#> <Training/Testing/Total>
#> <8/24/32>This is just scratching the surface of the new features and improvements in this release of rsample! You can see a full list of changes in the the release notes.
Acknowledgements
We’d like to thank everyone that has contributed since the last release: @DavisVaughan, @juliasilge, @mattwarkentin, @mikemahoney218, and @sametsoekel.
