The tidymodels framework is a collection of R packages for modeling and machine learning using tidyverse principles. There have been quite a number of updates and new developments in the tidymodels ecosystem since our last blog post in December! Since that post, tidymodels maintainers have published eight CRAN releases of existing packages. You can install these updates from CRAN with:
install.packages(c("broom", "butcher", "embed", "parsnip",
"rsample", "rules", "tune", "workflows"))
We purposefully write code in small, modular packages to make them easier to maintain (for us!) and use in production systems (for you!) but this does mean that sometimes any given package release can feel a bit minor. Some of the changes in these releases are small bug fixes or updates for changes in CRAN standards. However, there are also some substantively helpful new functions for modeling and resampling, and we want to make sure that folks can stay up-to-date with the changes and new features available.
We plan to begin regular updates every three or four months here on the tidyverse blog summarizing what’s happening lately in the tidymodels ecosystem overall. We’ll still continue the focused blog posts on more major new features that we’ve always written; look for one soon on a new package for creating and handling a collection of multiple modeling workflows all together. The NEWS files are linked here for each package, but read below for more details on some highlights that may interest you!
Choose parsnip models with an RStudio addin
The parsnip package provides support for a plethora of models. You can explore these models online at tidymodels.org, but the recent release of parsnip also contains an RStudio addin for choosing parsnip models and generating code to specify them.
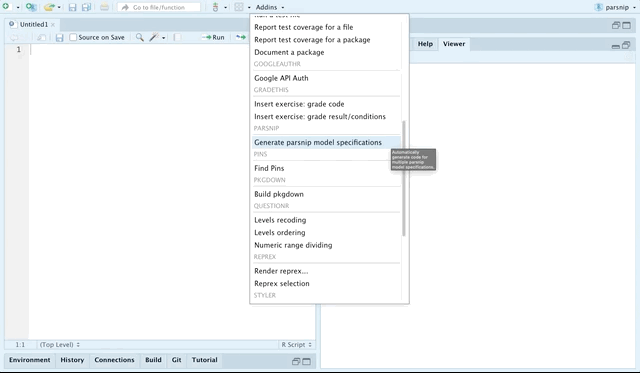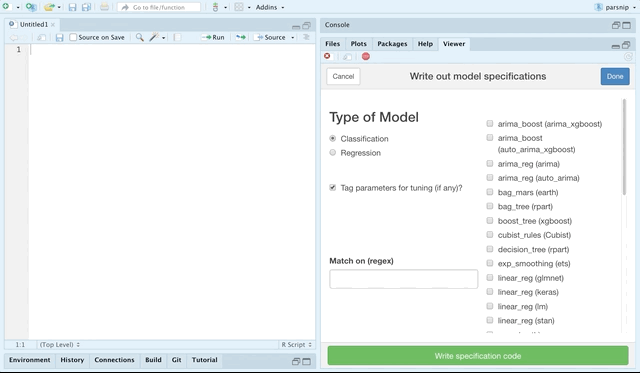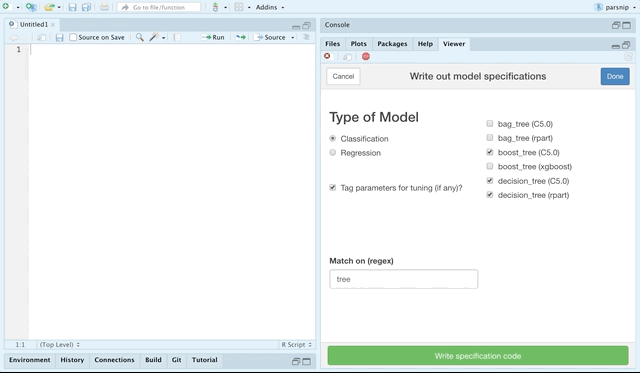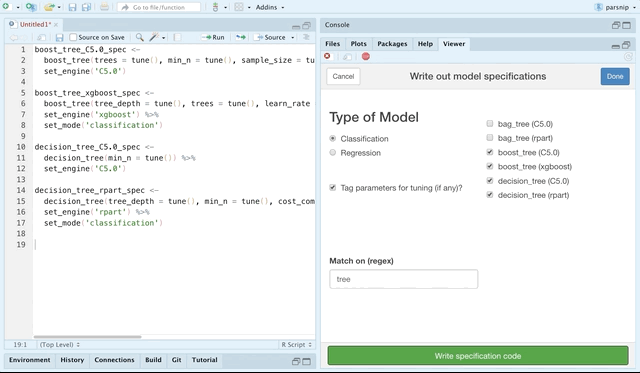
You can choose by classification or regression models, and even match by a regular expression.
There is now also
an augment() function for parsnip models, in addition to the augment() functions
for tuning results and
for workflows.
This recent screencast demonstrates how to use parsnip’s augment() function.
New functions in rsample
Most of the changes in the recent release for
rsample are internal and developer-facing, made to support rsample-adjacent packages like our new package for resampling spatial data (see below! 👀) but the new reg_intervals() function allows you to find bootstrap confidence intervals for simple models fluently. You have always been able to use rsample functions for
flexible bootstrap resampling but this new convenience function reduces the steps to get confidence intervals for models like lm() and glm().
library(rsample)
data(ad_data, package = "modeldata")
set.seed(123)
reg_intervals(
Class ~ tau + VEGF,
model_fn = "glm",
data = ad_data,
family = "binomial"
)
## # A tibble: 2 x 6
## term .lower .estimate .upper .alpha .method
## <chr> <dbl> <dbl> <dbl> <dbl> <chr>
## 1 tau -4.92 -4.11 -3.08 0.05 student-t
## 2 VEGF 0.651 0.959 1.22 0.05 student-t
Check out
my recent screencast for more details on using reg_intervals().
Also take a look at the
new permutations() function for permuting variables!
Resampling for spatial data
We are pleased to announce the first release of the spatialsample package.
You can install it from CRAN with:
install.packages("spatialsample")
The goal of spatialsample is to provide functions and classes for spatial resampling to use with
rsample. We intend to grow the number of spatial resampling approaches included in the package; the initial release includes spatial_clustering_cv(), a straightforward spatial resampling strategy with light dependencies based on k-means clustering.
library(spatialsample)
data("ames", package = "modeldata")
set.seed(234)
folds <- spatial_clustering_cv(ames, coords = c("Latitude", "Longitude"), v = 5)
folds
## # 5-fold spatial cross-validation
## # A tibble: 5 x 2
## splits id
## <list> <chr>
## 1 <split [2277/653]> Fold1
## 2 <split [2767/163]> Fold2
## 3 <split [2040/890]> Fold3
## 4 <split [2567/363]> Fold4
## 5 <split [2069/861]> Fold5
In this example, the ames data on houses in Ames, IA is resampled with v = 5; notice that the resulting partitions do not contain an equal number of observations.
We can create a helper plotting function to visualize the five folds.
library(ggplot2)
library(purrr)
library(dplyr)
plot_splits <- function(split) {
p <- analysis(split) %>%
mutate(analysis = "Analysis") %>%
bind_rows(assessment(split) %>%
mutate(analysis = "Assessment")) %>%
ggplot(aes(Longitude, Latitude, color = analysis)) +
geom_point(alpha = 0.5) +
labs(color = NULL)
print(p)
}
walk(folds$splits, plot_splits)
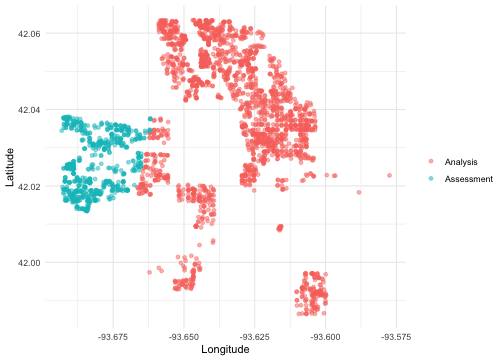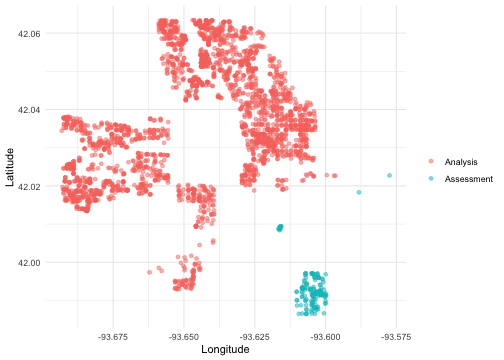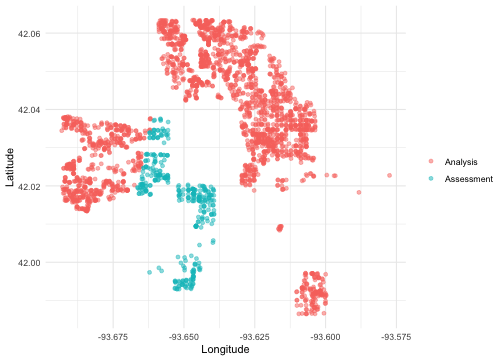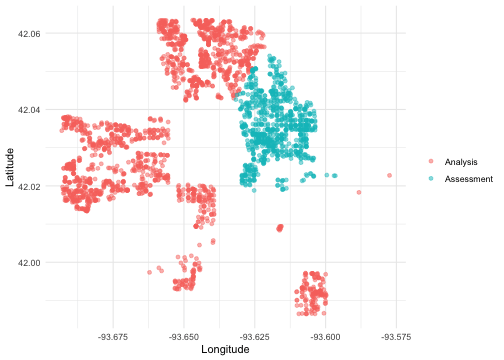
Check out the main vignette to see how this resampling strategy can be used for modeling, and submit an issue if there is a particular spatial resampling approach that you are interested in us prioritizing for future releases.
Acknowledgements
A big thanks to all of the contributors who helped make these releases possible! For some of these packages (like rsample, butcher, and embed), we have never said thank you before so we’ll take this opportunity to express our appreciation.
- broom: @AdroMine, @alexpghayes, @Amogh-Joshi, @anddis, @andrjohns, @AntoniosBarotsis, @arthur-e, @asreece, @asshah4, @briatte, @bwiernik, @cbhurley, @clausherther, @clauswilke, @crsh, @DarwinAwardWinner, @deblnia, @deschen1, @eheinzen, @friendly, @grantmcdermott, @hasandiwan, @hd-barros, @hfrick, @hughjonesd, @IndrajeetPatil, @irkaal, @jiho, @jmbarbone, @joshyam-k, @JReising09, @julian-urbano, @juliasilge, @kjfarley, @kristyrobledo, @leeweizhe1993, @leungi, @LukasWallrich, @matthieu-faron, @MatthieuStigler, @milanwiedemann, @mk9y, @mlatif71, @mlaviolet, @Nateme16, @nlubock, @pachamaltese, @rudeboybert, @saadaslam, @simonpcouch, @tavareshugo, @uqzwang, @vincentarelbundock, @WillemVervoort, and @zief0002
- butcher: @abichat, @adtserapio, @DavisVaughan, @edwinschut, @hfrick, @irkaal, @jarauh, @juliasilge, @jyuu, @kevinykuo, @klin333, @mkearney, @natejessee, @topepo, and @UnclAlDeveloper
- embed: @agilebean, @ajing, @Athospd, @Cardosaum, @ciberger, @data-datum, @dfalbel, @EmilHvitfeldt, @goleng, @hfrick, @ismailmuller, @juliasilge, @konradsemsch, @kylegilde, @lorenzwalthert, @mlduarte, @nhward, @niszet, @quantumlinguist, @smingerson, @tmastny, @tonigril, and @topepo
- parsnip: @awunderground, @Bijaelo, @DavisVaughan, @derek-corcoran-barrios, @eamoncaddigan, @EmilHvitfeldt, @ericpgreen, @hfrick, @irkaal, @jjcurtin, @joeycouse, @juliasilge, @kwiscion, @kylegilde, @lorenzwalthert, @markfairbanks, @mdancho84, @mlane3, @mrepetto94, @ndiquattro, @rorynolan, @shosaco, @smingerson, @tanho63, and @topepo
- rsample: @alexpghayes, @apreshill, @Athospd, @brunocarlin, @ColinConwell, @cportner, @danilinares, @DanOvando, @DavisVaughan, @dchiu911, @Dpananos, @dpastling, @EmilHvitfeldt, @fbchow, @fusaroli, @gcameron89777, @gregrs-uk, @gtalckmin, @hfrick, @hlynurhallgrims, @irkaal, @issactoast, @JamesM131, @johnaeanderson, @jonkeane, @juliasilge, @jyuu, @krlmlr, @kylegilde, @mattwarkentin, @mdancho84, @msmith01, @MxNl, @NikolaiVogl, @oude-gao, @PathosEthosLogos, @RMHogervorst, @sccmckenzie, @Shu-Wan, @skeller88, @skinnider, @sschooler, @swt30, @tjmahr, @tmastny, @topepo, and @UnclAlDeveloper
- rules: @frequena, @hfrick, @irkaal, @jaredlander, @juliasilge, @topepo, and @vidarsumo
- tune: @DavisVaughan, @HenrikBengtsson, @hfrick, @juliasilge, @kevin-m-kent, @kylegilde, @mine-cetinkaya-rundel, @rorynolan, @siegfried, @stevenpawley, and @topepo
