I’m very excited to announce the release of tidyr 1.0.0! tidyr provides a set of tools for transforming data frames to and from tidy data, where each variable is a column and each observation is a row. Tidy data is a convention for matching the semantics and structure of your data that makes using the rest of the tidyverse (and many other R packages) much easier.
Install tidyr with:
install.packages("tidyr")
As you might guess from the version number, this is a major release, and the 1.0.0 moniker indicates that I’m finally happy with the overall interface of the package. This has been a long time coming: it’s five years since the first tidyr release, nine years since the first reshape2 release, and fourteen years since the first reshape release!
This blog post summarises the four major changes to the package:
New
pivot_longer()andpivot_wider()provide improved tools for reshaping, supercedingspread()andgather(). The new functions are substantially more powerful, thanks to ideas from the data.table and cdata packages, and I’m confident that you’ll find them easier to use and remember than their predecessors.New
unnest_auto(),unnest_longer(),unnest_wider(), andhoist()provide new tools for rectangling, converting deeply nested lists into tidy data frames.nest()andunnest()have been changed to match an emerging principle for the design of...interfaces. Four new functions (pack()/unpack(), andchop()/unchop()) reveal that nesting is the combination of two simpler steps.New
expand_grid(), a variant ofbase::expand.grid(). This is a useful function to know about, but also serves as a good reason to discuss the important role that vctrs plays behind the scenes. You shouldn’t ever have to learn about vctrs, but it brings improvements to consistency and performance.
As well as implementing the new features, I’ve spent considerable time on the documentation, including four major new vignettes:
vignette("pivot"),vignette("rectangle"), andvignette("nest")provide detailed documentation and case studies of pivotting, rectangling, and nesting respectively.vignette("in-packages")provides best practices for using tidyr inside another package, and detailed advice on working with multiple versions of tidyr if an interface change has affected your package.
You can see a list of all the other minor bug fixes and improvements in the release notes. I strongly recommend reading the complete release notes if you’re a package developer.
library(tidyr)
library(dplyr)
Pivoting
New
pivot_longer() and
pivot_wider() provide modern alternatives to spread() and gather(). They have been carefully redesigned to be easier to learn and remember, and include many new features. spread() and gather() won’t go away, but they’ve been retired which means that they’re no longer under active development.
The best place to learn about pivot_longer() and pivot_wider() is
vignette("pivot"), or by watching my presentation to the
Vienna R users group. Here I’ll quickly show off a few of the coolest new features:
pivot_longer()can now separate column names into multiple variables in a single step. For example, take thewhodataset which has column names that look likenew_{diagnosis}_{gender}{age}:names(who) #> [1] "country" "iso2" "iso3" "year" #> [5] "new_sp_m014" "new_sp_m1524" "new_sp_m2534" "new_sp_m3544" #> [9] "new_sp_m4554" "new_sp_m5564" "new_sp_m65" "new_sp_f014" #> [13] "new_sp_f1524" "new_sp_f2534" "new_sp_f3544" "new_sp_f4554" #> [17] "new_sp_f5564" "new_sp_f65" "new_sn_m014" "new_sn_m1524" #> [21] "new_sn_m2534" "new_sn_m3544" "new_sn_m4554" "new_sn_m5564" #> [25] "new_sn_m65" "new_sn_f014" "new_sn_f1524" "new_sn_f2534" #> [29] "new_sn_f3544" "new_sn_f4554" "new_sn_f5564" "new_sn_f65" #> [33] "new_ep_m014" "new_ep_m1524" "new_ep_m2534" "new_ep_m3544" #> [37] "new_ep_m4554" "new_ep_m5564" "new_ep_m65" "new_ep_f014" #> [41] "new_ep_f1524" "new_ep_f2534" "new_ep_f3544" "new_ep_f4554" #> [45] "new_ep_f5564" "new_ep_f65" "newrel_m014" "newrel_m1524" #> [49] "newrel_m2534" "newrel_m3544" "newrel_m4554" "newrel_m5564" #> [53] "newrel_m65" "newrel_f014" "newrel_f1524" "newrel_f2534" #> [57] "newrel_f3544" "newrel_f4554" "newrel_f5564" "newrel_f65"You can now tease apart the variable names in a single step (i.e. without
separate())) by supplying a vector of variable names tonames_toand a regular expression tonames_pattern(simpler cases might only neednames_sep):who %>% pivot_longer( cols = new_sp_m014:newrel_f65, names_to = c("diagnosis", "gender", "age"), names_pattern = "new_?(.*)_(.)(.*)", values_to = "count", values_drop_na = TRUE ) #> # A tibble: 76,046 x 8 #> country iso2 iso3 year diagnosis gender age count #> <chr> <chr> <chr> <int> <chr> <chr> <chr> <int> #> 1 Afghanistan AF AFG 1997 sp m 014 0 #> 2 Afghanistan AF AFG 1997 sp m 1524 10 #> 3 Afghanistan AF AFG 1997 sp m 2534 6 #> 4 Afghanistan AF AFG 1997 sp m 3544 3 #> 5 Afghanistan AF AFG 1997 sp m 4554 5 #> 6 Afghanistan AF AFG 1997 sp m 5564 2 #> # … with 7.604e+04 more rowspivot_longer()can now work with rows that contain multiple observations (this feature was inspired by data.table’sdcast()method). For example, take the baseanscombedataset. Each row consists of four pairs ofxandymeasurements:head(anscombe) #> x1 x2 x3 x4 y1 y2 y3 y4 #> 1 10 10 10 8 8.04 9.14 7.46 6.58 #> 2 8 8 8 8 6.95 8.14 6.77 5.76 #> 3 13 13 13 8 7.58 8.74 12.74 7.71 #> 4 9 9 9 8 8.81 8.77 7.11 8.84 #> 5 11 11 11 8 8.33 9.26 7.81 8.47 #> 6 14 14 14 8 9.96 8.10 8.84 7.04We can now tidy this in a single step by using the special
.valuevariable name:anscombe %>% pivot_longer( everything(), names_to = c(".value", "set"), names_pattern = "(.)(.)" ) %>% as_tibble() #> # A tibble: 44 x 3 #> set x y #> <chr> <dbl> <dbl> #> 1 1 10 8.04 #> 2 2 10 9.14 #> 3 3 10 7.46 #> 4 4 8 6.58 #> 5 1 8 6.95 #> 6 2 8 8.14 #> # … with 38 more rowspivot_wider()can now do simple aggregations (reshape2::dcast()fans rejoice!). For example, take the basewarpbreaksdataset (converted to a tibble to print more compactly):warpbreaks <- warpbreaks %>% as_tibble() %>% select(wool, tension, breaks) warpbreaks #> # A tibble: 54 x 3 #> wool tension breaks #> <fct> <fct> <dbl> #> 1 A L 26 #> 2 A L 30 #> 3 A L 54 #> 4 A L 25 #> 5 A L 70 #> 6 A L 52 #> # … with 48 more rowsThis is a designed experiment with nine replicates for every combination of
wool(AandB) andtension(L,M,H). If we attempt to pivot the levels ofwoolinto the columns, we get a warning and the output contains list-columns:warpbreaks %>% pivot_wider(names_from = wool, values_from = breaks) #> Warning: Values in `breaks` are not uniquely identified; output will contain list-cols. #> * Use `values_fn = list(breaks = list)` to suppress this warning. #> * Use `values_fn = list(breaks = length)` to identify where the duplicates arise #> * Use `values_fn = list(breaks = summary_fun)` to summarise duplicates #> # A tibble: 3 x 3 #> tension A B #> <fct> <list<dbl>> <list<dbl>> #> 1 L [9] [9] #> 2 M [9] [9] #> 3 H [9] [9]You can now summarise the duplicates with the
values_fnargument:warpbreaks %>% pivot_wider( names_from = wool, values_from = breaks, values_fn = list(breaks = mean) ) #> # A tibble: 3 x 3 #> tension A B #> <fct> <dbl> <dbl> #> 1 L 44.6 28.2 #> 2 M 24 28.8 #> 3 H 24.6 18.8
Learn the full details and see many more examples in
vignette("pivot").
Rectangling
Rectangling is the art and craft of taking a deeply nested list (often sourced from wild-caught JSON or XML) and taming it into a tidy data set of rows and columns. tidyr 1.0.0 provides four new functions to aid rectangling:
unnest_longer()takes each element of a list-column and makes a new row.unnest_wider()takes each element of a list-column and makes a new column.unnest_auto()uses some heuristics to guess whether you wantunnest_longer()orunnest_wider().hoist()is similar tounnest_wider()but only plucks out selected components, and can reach down multiple levels.
To see them in action, take this small sample from repurrrsive::got_chars. It contains data about three characters from the Game of Thrones:
characters <- list(
list(
name = "Theon Greyjoy",
aliases = c("Prince of Fools", "Theon Turncloak", "Theon Kinslayer"),
alive = TRUE
),
list(
name = "Tyrion Lannister",
aliases = c("The Imp", "Halfman", "Giant of Lannister"),
alive = TRUE
),
list(
name = "Arys Oakheart",
alive = FALSE
)
)
To work with the new tidyr rectangling tools, we first put the list into a data frame, creating a list-column:
got <- tibble(character = characters)
got
#> # A tibble: 3 x 1
#> character
#> <list>
#> 1 <named list [3]>
#> 2 <named list [3]>
#> 3 <named list [2]>
We can then use unnest_wider() to make each element of that list into a column:
got %>%
unnest_wider(character)
#> # A tibble: 3 x 3
#> name aliases alive
#> <chr> <list> <lgl>
#> 1 Theon Greyjoy <chr [3]> TRUE
#> 2 Tyrion Lannister <chr [3]> TRUE
#> 3 Arys Oakheart <???> FALSE
Followed by unnest_longer() to turn each alias into its own row:
got %>%
unnest_wider(character) %>%
unnest_longer(aliases)
#> # A tibble: 7 x 3
#> name aliases alive
#> <chr> <chr> <lgl>
#> 1 Theon Greyjoy Prince of Fools TRUE
#> 2 Theon Greyjoy Theon Turncloak TRUE
#> 3 Theon Greyjoy Theon Kinslayer TRUE
#> 4 Tyrion Lannister The Imp TRUE
#> 5 Tyrion Lannister Halfman TRUE
#> 6 Tyrion Lannister Giant of Lannister TRUE
#> 7 Arys Oakheart <NA> FALSE
Even more conveniently, you can use unnest_auto() to guess which direction a list column should be unnested in. Here it yields the same results as above, and the messages tell you why:
got %>%
unnest_auto(character) %>%
unnest_auto(aliases)
#> Using `unnest_wider(character)`; elements have 2 names in common
#> Using `unnest_longer(aliases)`; no element has names
Alternatively, you can use hoist() to reach deeply into a data structure and put out just the pieces you need:
got %>% hoist(character,
name = "name",
alias = list("aliases", 1),
alive = "alive"
)
#> # A tibble: 3 x 4
#> name alias alive character
#> <chr> <chr> <lgl> <list>
#> 1 Theon Greyjoy Prince of Fools TRUE <named list [1]>
#> 2 Tyrion Lannister The Imp TRUE <named list [1]>
#> 3 Arys Oakheart <NA> FALSE <named list [0]>
This syntax provides a more approachable alternative to using purrr::map() inside dplyr::mutate(), as we’d previously recommended:
got %>% mutate(
name = purrr::map_chr(character, "name"),
alias = purrr::map_chr(character, list("aliases", 1), .default = NA),
alive = purrr::map_lgl(character, "alive")
)
Learn more in
vignette("rectangle").
Nesting
nest() and unnest() have been updated with new interfaces that are more closely aligned to evolving tidyverse conventions. The biggest change is to their operation with multiple columns:
# old
df %>% nest(x, y, z)
# new
df %>% nest(data = c(x, y, z))
# old
df %>% unnest(x, y, z)
# new
df %>% unnest(c(x, y, z))
I’ve done my best to ensure that common uses of nest() and unnest() will continue to work, generating an informative warning telling you precisely how you need to update your code. If this doesn’t work, you can use nest_legacy() or unnest_legacy() to access the previous interfaces; see
vignette("in-packages") for more advice on managing this transition.
Behind the scenes, I discovered that nesting (and unnesting) can be decomposed into the combination of two simpler operations:
pack()andunpack()multiple columns into and out of a data frame column.chop()andunchop()chop up rows into and out of list-columns. It’s a bit like an explicit form ofdplyr::group_by().
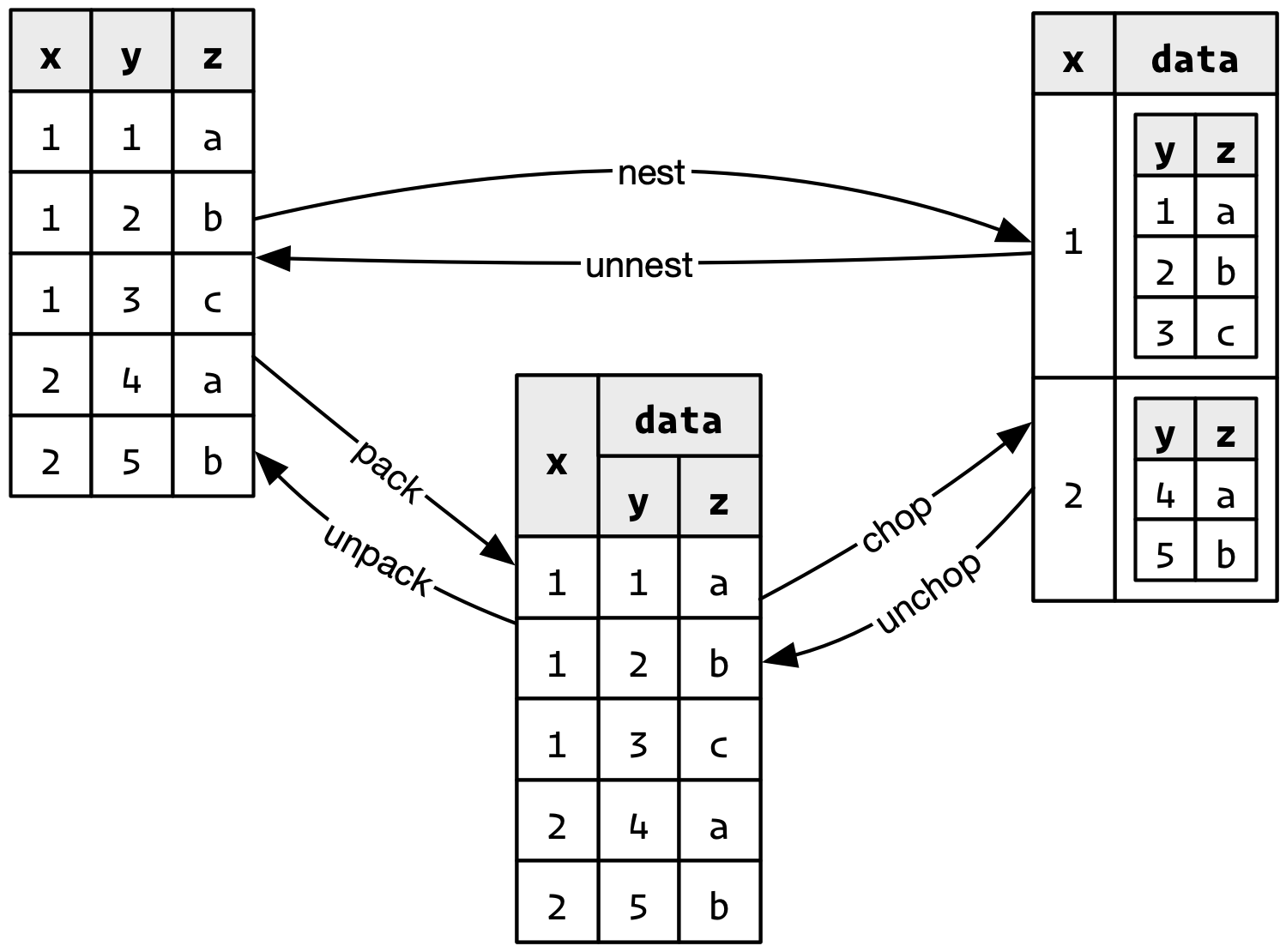
This is primarily of internal interest, but it considerably simplifies the implementation of nest(), and you may occasionally find the underlying functions useful when working with exotic data structures.
expand_grid()
expand_grid() completes the existing family of expand(), nesting(), and crossing() with a low-level function that works with vectors:
expand_grid(
x = 1:3,
y = letters[1:3],
z = LETTERS[1:3]
)
#> # A tibble: 27 x 3
#> x y z
#> <int> <chr> <chr>
#> 1 1 a A
#> 2 1 a B
#> 3 1 a C
#> 4 1 b A
#> 5 1 b B
#> 6 1 b C
#> # … with 21 more rows
Compared to the existing base function expand.grid(), expand_grid():
- Varies the first element most slowly (not most quickly).
- Never converts strings to factors and doesn’t add any additional attributes.
- Returns a tibble, not a data frame.
- Can expand data frames.
The last feature is quite powerful, as it allows you to generate partial grids:
students <- tribble(
~ school, ~ student,
"A", "John",
"A", "Mary",
"A", "Susan",
"B", "John"
)
expand_grid(students, semester = 1:2)
#> # A tibble: 8 x 3
#> school student semester
#> <chr> <chr> <int>
#> 1 A John 1
#> 2 A John 2
#> 3 A Mary 1
#> 4 A Mary 2
#> 5 A Susan 1
#> 6 A Susan 2
#> 7 B John 1
#> 8 B John 2
This is made possible by the vctrs package. vctrs is primarily of interest to package developers but I want to talk about it briefly here because I’ve been having a lot of fun working with it. It’s hard to concisely describe vctrs, but one aspect is carefully defining what a “vector” is, and providing a set of useful functions that work on all types of vctrs, without any special cases. One interesting finding is that thinking of a data frame as a vector of rows (not columns, as R usually does) is suprisingly useful, and something you can expect to see in more places in the tidyverse in the future.
Note that when data frame inputs are unnamed, they’re automatically unpacked into individual columns in the output. It’s also possible to create a column that is itself a data frame, a df-column, if you name it:
expand_grid(student = students, semester = 1:2)
#> # A tibble: 8 x 2
#> student$school $student semester
#> <chr> <chr> <int>
#> 1 A John 1
#> 2 A John 2
#> 3 A Mary 1
#> 4 A Mary 2
#> 5 A Susan 1
#> 6 A Susan 2
#> 7 B John 1
#> 8 B John 2
Df-columns aren’t particularly useful yet, but they provide powerful building blocks for future extensions. For example, we expect a future version of dplyr will support df-columns as a way for mutate() and summarise() to create multiple new columns from a single function call.
Thanks
A big thanks to all 95 people who help to make this release possible! @abiyug, @AdvikS, @ahcyip, @alexpghayes, @aneisse, @apreshill, @atusy, @ax42, @banfai, @billdenney, @brentthorne, @brunj7, @coolbutuseless, @courtiol, @cwickham, @DanielReedOcean, @DavisVaughan, @dewittpe, @donboyd5, @earowang, @eElor, @enricoferrero, @fresques, @garrettgman, @gederajeg, @georgevbsantiago, @giocomai, @gireeshkbogu, @gorkang, @ha0ye, @hadley, @hplieninger, @iago-pssjd, @IndrajeetPatil, @jackdolgin, @japhir, @jayhesselberth, @jennybc, @jeroenjanssens, @jestarr, @jgendrinal, @Jim89, @jl5000, @jmcastagnetto, @justasmundeikis, @Kaz272, @KenatRSF, @koncina, @krlmlr, @kuriwaki, @lazappi, @leeper, @lionel-, @markdly, @martinjhnhadley, @MatthieuStigler, @mattwarkentin, @mehrgoltiv, @meriops, @mikemc, @mikmart, @mine-cetinkaya-rundel, @mitchelloharawild, @mixolydianpink, @mkapplebee, @moodymudskipper, @mpaulacaldas, @Myfanwy, @ogorodriguez, @onesandzeroes, @Overlytic, @paleolimbot, @PMassicotte, @psychometrician, @Rekyt, @romainfrancois, @romatik, @rubenarslan, @SchollJ, @seabbs, @sethmund, @sfirke, @SimonCoulombe, @Stephonomon, @stufield, @tdjames1, @thierrygosselin, @tklebel, @tmastny, @trannhatanh89, @TrentLobdell, @wcmbishop, @wulixin, @yutannihilation, and @zeehio.
