We’re excited to announce version 1.4.1 of the tibble package. Tibbles are a modern reimagining of the data frame, keeping what time has shown to be effective, and throwing out what is not, with nicer default output too! Grab the latest version with:
install.packages("tibble")This release contains the following major changes:
- Colored terminal output
- Compatibility fixes for code that expects data frames
- Improved
add_column()
There are many other small improvements and bug fixes: please see the release notes for a complete list.
Thanks to Anh Le for the add_cases() alias, to Davis Vaughan for improving add_column(), to Patrick O. Perry for converting C++ code to C, and to all the other contributors. Use the issue tracker to submit bugs or suggest ideas, your contributions are always welcome.
Colored terminal output
The most important change of this release is the use of color and other markup for the output in the terminal:
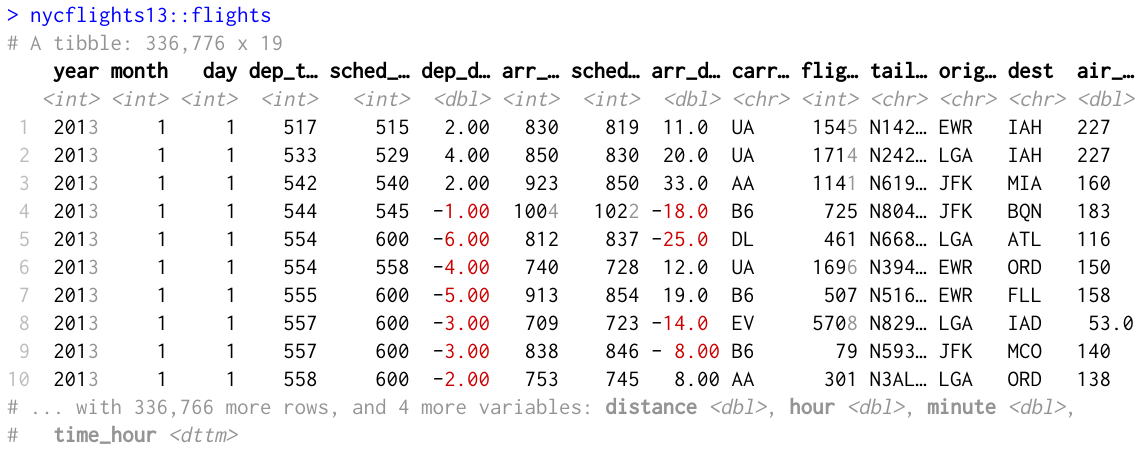
Numeric columns have two displays. If there is room, the full number is displayed, making it easy to compare order of magnitude. Digits after the first three are dimmed to emphasise the important components. Negative numbers are shown in red. The display switches to scientific format if there is not enough room.
Missing values get a yellow background. This makes it easier to distinguish the string
"NA"from a missing value.Long character values and factors are abbreviated in order to fit more columns on screen.
The highlighting is implemented by the new pillar package, which also offers extension points for packages such as hms that provide custom data types. See the Extending tibble vignette for details. We’re still figuring out how to use colour most effectively, so expect this to change in minor ways in the in future. If you have feedback, please let us know by filing an issue in the pillar package.
Compatibility fixes
Subsetting with logical or character indexes is now supported, just like with regular data frames:
tbl <- tibble(a = 1:3, b = letters[1:3])
tbl
#> # A tibble: 3 x 2
#> a b
#> <int> <chr>
#> 1 1 a
#> 2 2 b
#> 3 3 c
tbl[c(TRUE, FALSE, TRUE), ]
#> # A tibble: 2 x 2
#> a b
#> <int> <chr>
#> 1 1 a
#> 2 3 c
tbl[c("2", "odd-row-name"), ]
#> # A tibble: 2 x 2
#> a b
#> <int> <chr>
#> 1 2 b
#> 2 NA <NA>Passing drop = TRUE is now supported and gives the same result as for regular data frames:
tbl[1, 2, drop = TRUE]
#> [1] "a"
tbl[1, drop = TRUE]
#> Warning: drop ignored
#> # A tibble: 3 x 1
#> a
#> <int>
#> 1 1
#> 2 2
#> 3 3Both changes make it easier to use tibbles with code that is designed to work with data frames.
Improved addition of columns
The add_column() functions now keeps all attributes of the original tibble, this is important for packages that extend tibbles such as sf or tibbletime.
library(tibbletime)
#>
#> Attaching package: 'tibbletime'
#> The following object is masked from 'package:stats':
#>
#> filter
tbl_with_date <-
tibble(a = 1:3, date = Sys.Date() + a) %>%
tbl_time(date)
add_column(tbl_with_date, b = letters[1:3])
#> # A time tibble: 3 x 3
#> # Index: date
#> a date b
#> * <int> <date> <chr>
#> 1 1 2018-01-04 a
#> 2 2 2018-01-05 b
#> 3 3 2018-01-06 cTrailing comma
You can now add a trailing comma in function calls with an ... argument:
tibble(
a = 1:3,
b = letters[1:3],
)
#> # A tibble: 3 x 2
#> a b
#> <int> <chr>
#> 1 1 a
#> 2 2 b
#> 3 3 c
add_column(
tbl,
c = LETTERS[1:3],
)
#> # A tibble: 3 x 3
#> a b c
#> <int> <chr> <chr>
#> 1 1 a A
#> 2 2 b B
#> 3 3 c CThis simplifies extending or adapting the code later, because you don’t need to remember to add or remove the trailing comma of the last function argument. (This functionality is available for all packages that support tidy evaluation, rlang >= 0.1.6 is required.)
Acknowledgments
We received issues, pull requests, and comments from 109 people since tibble 1.2.0. Thanks to everyone: @aalexandersson, @adnbps, @AkhilNairAmey, @alexhallam, @alibat, @amjiuzi, @AndreMikulec, @andrewjpfeiffer, @ashiklom, @atribe, @bapfeld, @barnettjacob, @behrman, @BillDunlap, @BruceZhaoR, @cassiusoat, @cboettig, @cderv, @ckluss, @ClaytonJY, @colearendt, @csgillespie, @dalejbarr, @dan87134, @DavisVaughan, @ddiez, @dhicks, @dlpd, @drewgendreau, @drolejoel, @echasnovski, @edzer, @ElsLommelen, @etiennebr, @FabianRoger, @garrettgman, @gavinsimpson, @geotheory, @ginolhac, @hadley, @happyshows, @heavywatal, @helix123, @holstius, @huftis, @ianmcook, @imanuelcostigan, @janschulz, @javierluraschi, @jennybc, @jimhester, @joelgombin, @jonathan-g, @kendonB, @kevinushey, @khughitt, @kismsu, @krlmlr, @kwstat, @LaDilettante, @lcolladotor, @lionel-, @lpmarco, @m-sostero, @MarcusWalz, @matteodefelice, @mattfidler, @mgirlich, @michaellevy, @MikeBadescu, @mkearney, @mmuurr, @Monduiz, @mubeenarasack, @mundl, @nbenn, @ncarchedi, @NikNakk, @noamross, @ntguardian, @p0bs, @patperry, @pgensler, @phalexo, @pssguy, @r2evans, @rentrop, @richierocks, @Rongpeng, @s-fleck, @sainathadapa, @sebschub, @sibojan, @slonik-az, @sskim47, @t-kalinowski, @thercast, @thornend, @tjmahr, @trinker, @vnijs, @vspinu, @vvrably, @wibom, @wpetry, @yeedle, @yihui, @yutannihilation, and @Zedseayou.
